Writing an R package interface
to C++ libraries
with Rcpp
@heavywatal
 @heavywatal
@heavywatal
 Watal M. Iwasaki = 岩嵜 航
Watal M. Iwasaki = 岩嵜 航https://heavywatal.github.io/
- PhD in Life Sciences, Tohoku University, Sendai
- Evolutionary theory of complexity and diversity in biological systems.
- Postdoc in SOKENDAI, Hayama
- Evolution of diversity within a tumor/cancer.

- Likes
- 🍺 Beer, Sake, Whisky, Cooking
- ♬ Heavy Metal, Classical, Folk
tumopp — tumor growth simulator in C++
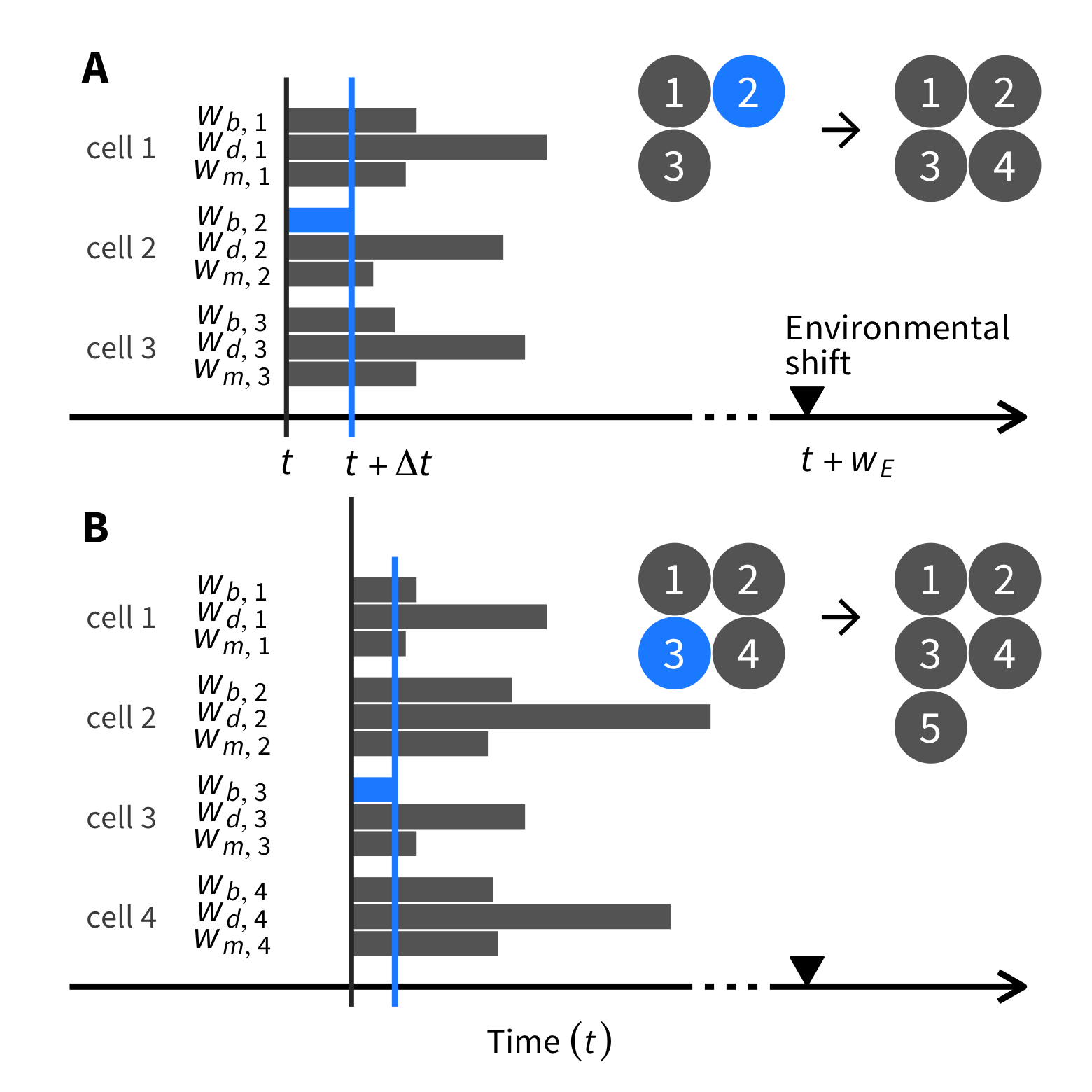
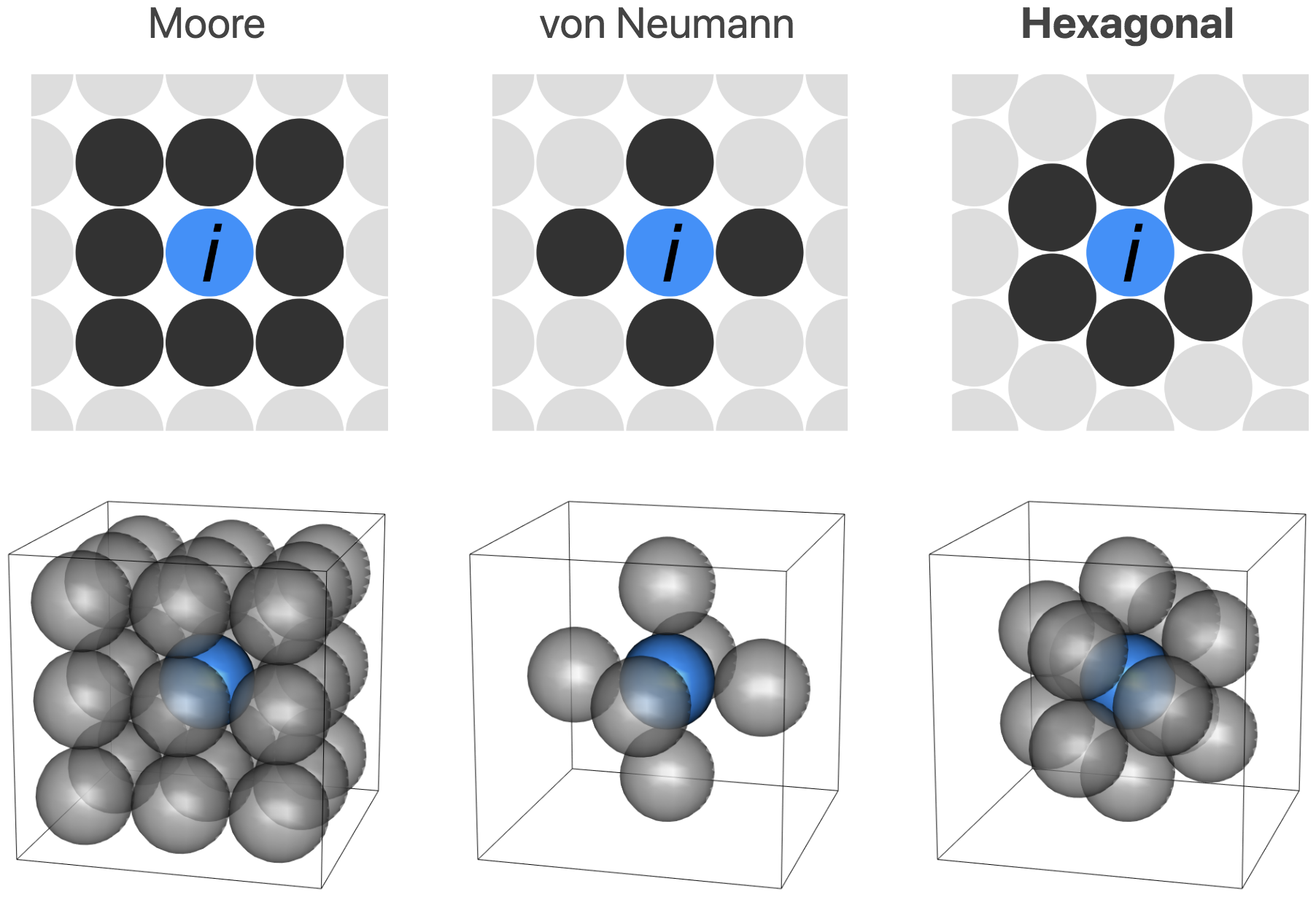
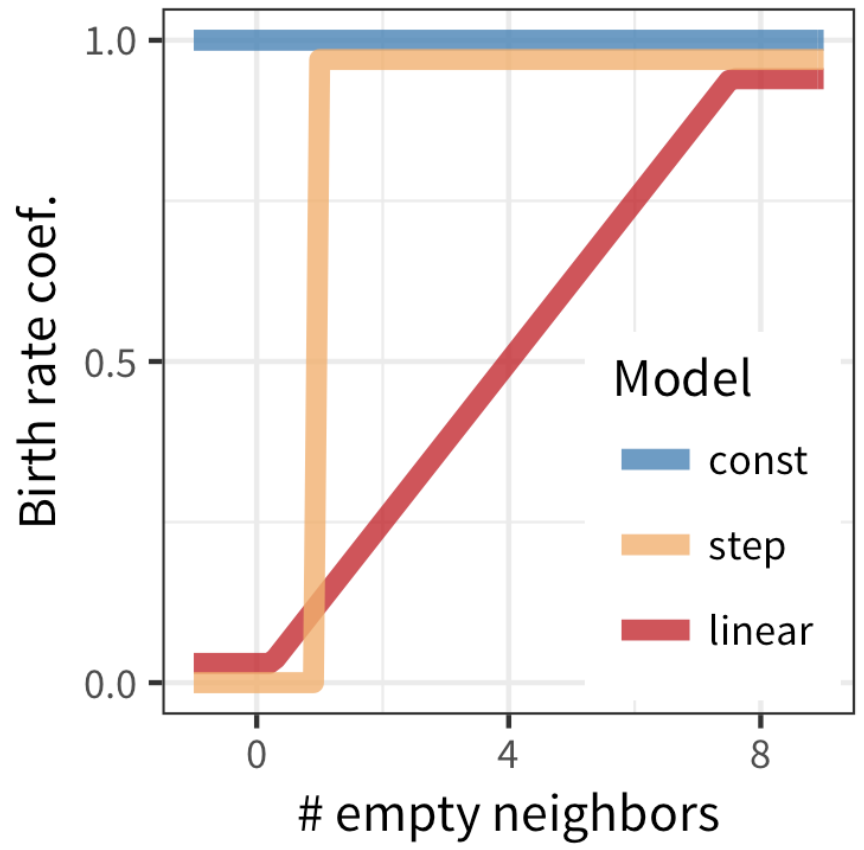
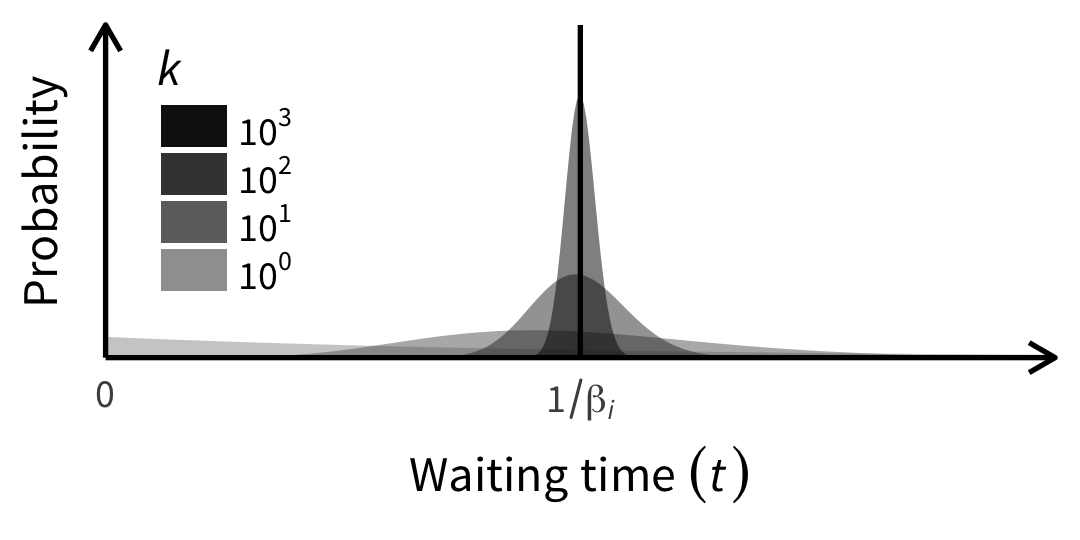
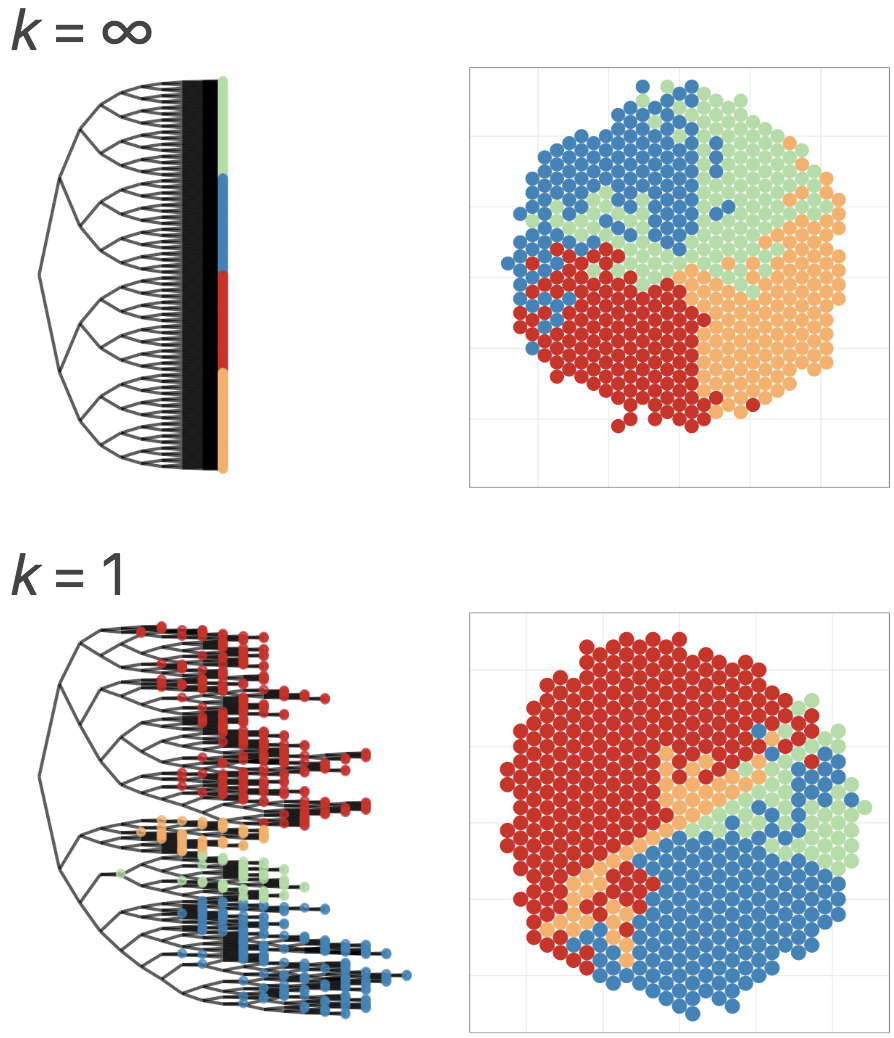
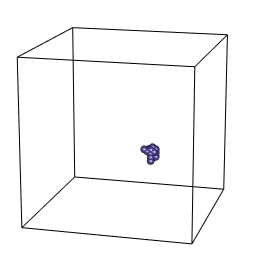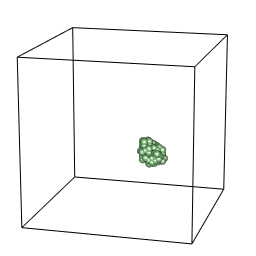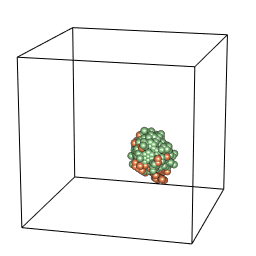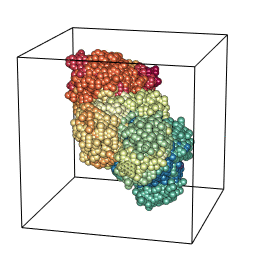

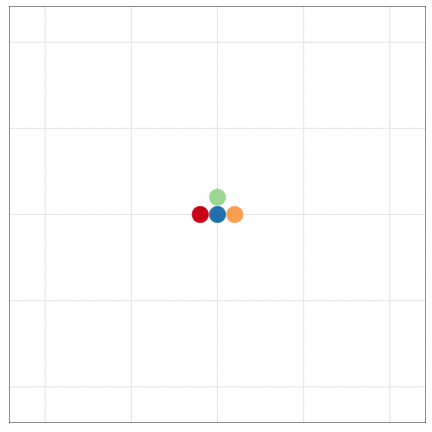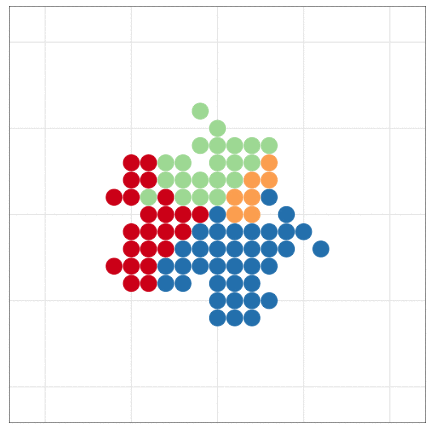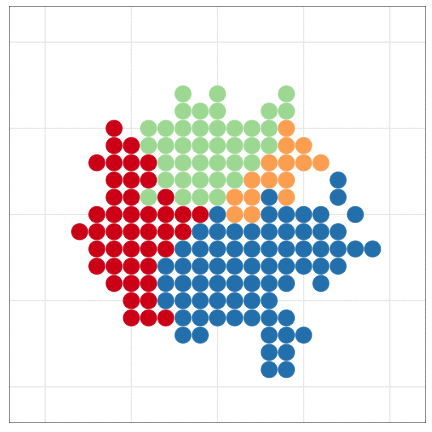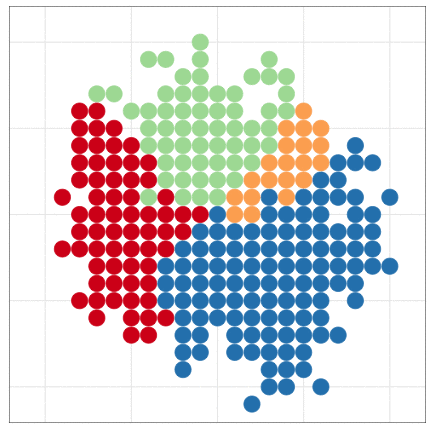
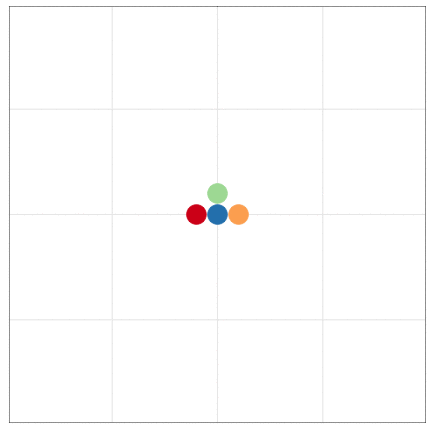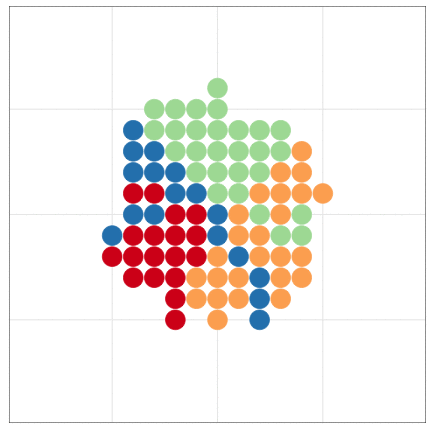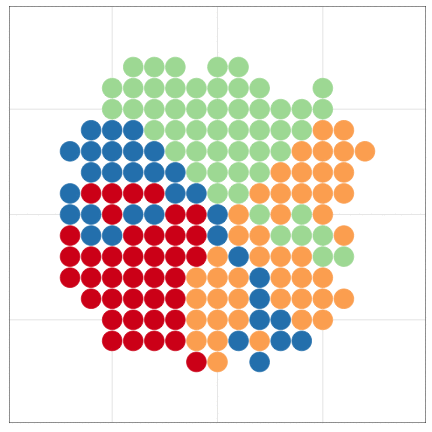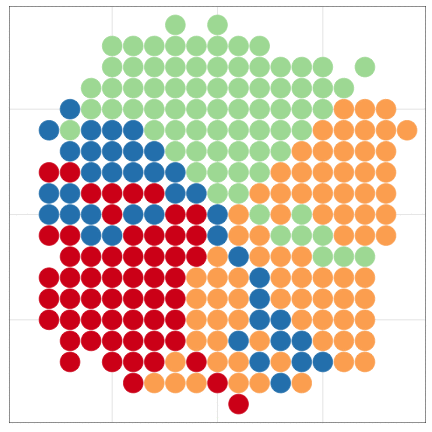
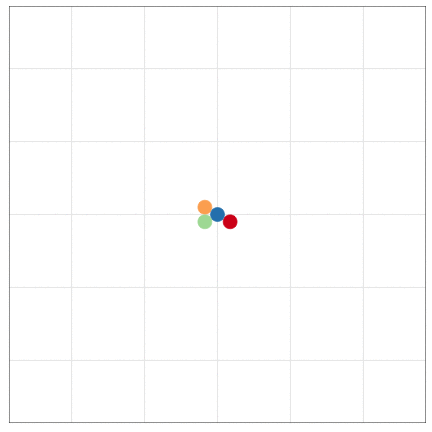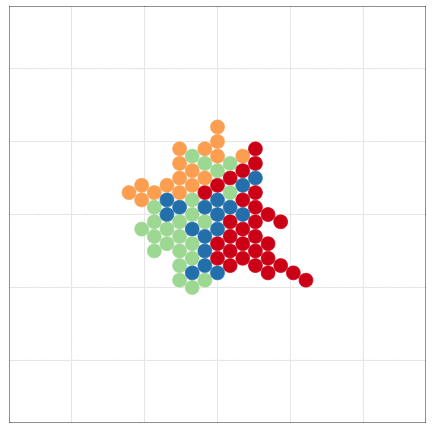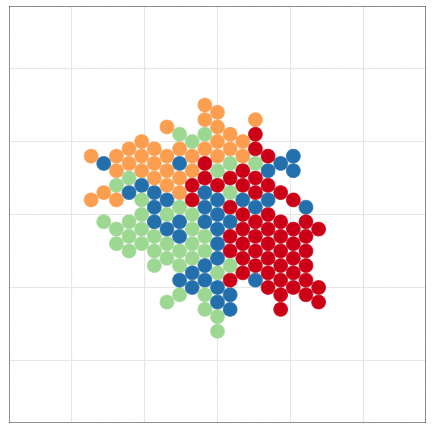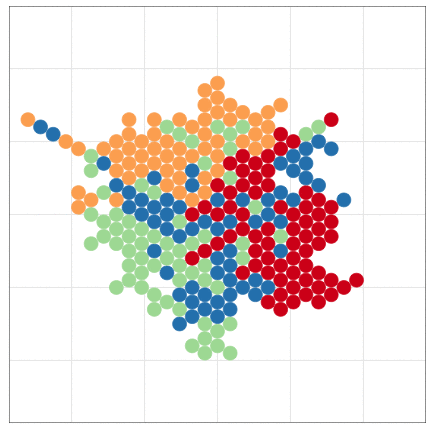
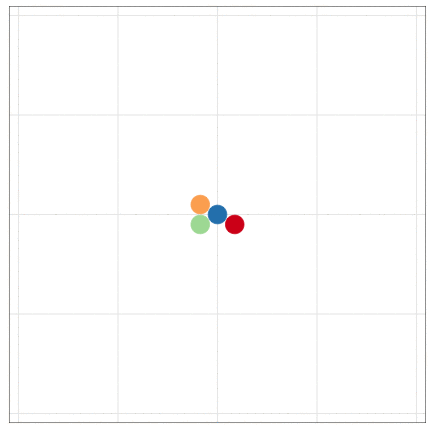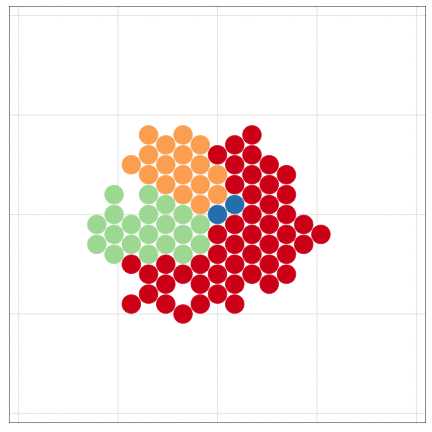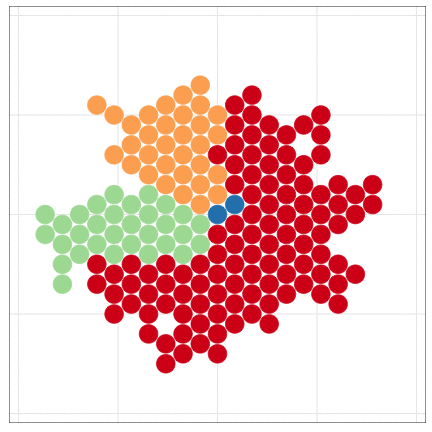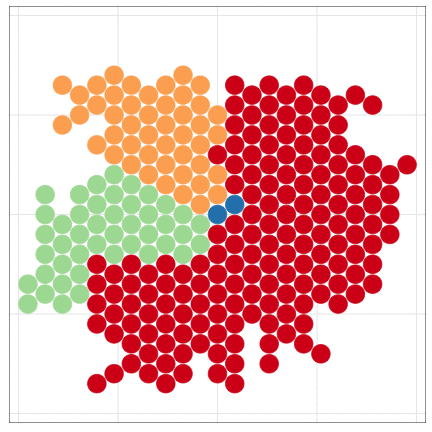
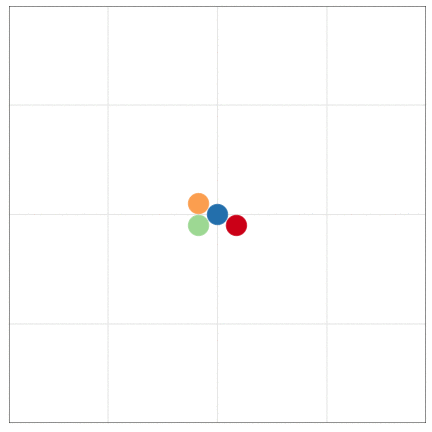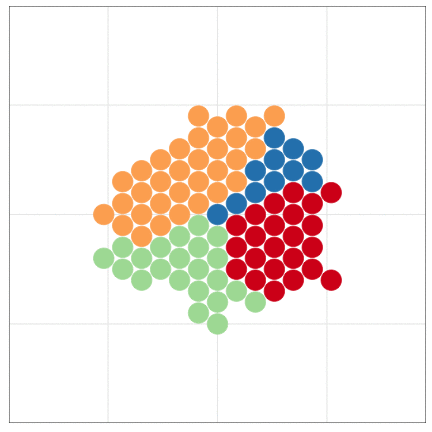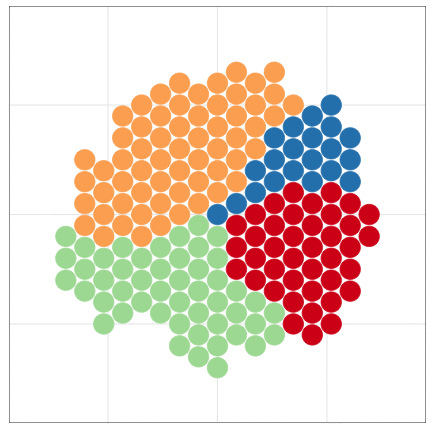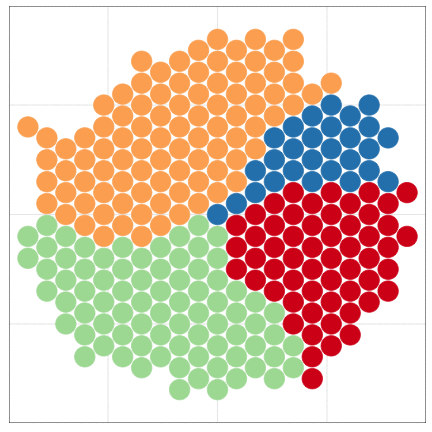
tumopp — tumor growth simulator in C++
Available via Homebrew/Linuxbrew:
brew install heavywatal/tap/tumopp
tumopp -N10000 -D3 -Chex -k100 -d0.1 -m0.5 -o OUTPUT_DIR
Dependencies:
- Unix-like OS (macOS, Linux, etc.)
- C++14 compiler (clang++ >= Apple LLVM 8.1, g++ >= 5.3)
- CMake (>= 3.1)
- Boost C++ Libraries (>= 1.65.0)
- sfmt-class
- cxxwtl
tumopp — tumor growth simulator in C++
Library structure:
CMAKE_INSTALL_PREFIX/
├── bin/
│ └── tumopp
├── include/
│ └── tumopp/
│ ├── cell.hpp
│ ├── simulation.hpp
│ └── tissue.hpp
└── lib/
└── libtumopp.dylib # libtumopp.so on Linux
Output structure:
OUTPUT_DIR/
├── drivers.tsv.gz
├── population.tsv.gz
├── program_options.conf
└── snapshots.tsv.gz
Workflow
- Run tumopp with some parameter sets from the command line.
- Write results to TSV files
- Start R
- Read TSV files as data.frames
- Visualize and analyze with tidyverse packages
Not too bad.
But it will be more convenient if I can run tumopp in R:
library(tumopp)
results = tumopp(some_parameter_sets) %>% print()
max coord dimensions shape delta0 rho0 population drivers
<int> <char> <int> <int> <num> <num> <list> <list>
1: 10000 hex 3 100 0.1 0.5 <tbl_df> <tbl_df>
Seamless R and C++ integration with Rcpp
It is typically used to eliminate bottlenecks in R code.
Many online examples show the ways to define a short function:
library(Rcpp)
Rcpp::cppFunction('
int fibonacci(int x) {
if (x < 2) return x;
return fibonacci(x - 1) + fibonacci(x - 2);
}
')
fibonacci(8L)
[1] 21
But I already have C++ functions in my library.
How can I use them in R?
Create an R+Rcpp package from scratch
Package components (See http://r-pkgs.had.co.nz/ for details):
DESCRIPTION # Package metadata
LICENSE
NAMESPACE # List of objects to import/export
R/ # R code (*.R)
man/ # Object documentation (*.Rd)
src/ # C++ source code (*.cpp)
vignettes/ # Long-form guide
Use devtools and/or usethis to setup a skeleton:
usethis::create_package("tumopp")
usethis::use_mit_license()
usethis::use_roxygen_md()
usethis::use_package_doc()
usethis::use_rcpp()
usethis::use_git()
Then, modify DESCRIPTION, R/, and src/.
Package-wide settings in R/tumopp-package.R
#' @useDynLib tumopp, .registration = TRUE
#' @importFrom Rcpp sourceCpp
#' @importFrom magrittr %>%
#' @aliases NULL tumopp-package
#' @keywords internal
"_PACKAGE"
.onUnload = function(libpath) {
library.dynam.unload("tumopp", libpath)
}
@useDynLibis needed to import compiled C++ functions.@importFrom Rcpp sourceCppseems necessary to load Rcpp."_PACKAGE"is a special string to generate package documentation..onUnloadis recommended in http://adv-r.had.co.nz/Rcpp.html,
but not used in the major packages these days…?
Hadley said: “It is polite to define it, but easy to forget.”
Define Rcpp function to use external libraries
Create src/run.cpp:
// [[Rcpp::plugins(cpp14)]]
#include <Rcpp.h>
#include <tumopp/simulation.hpp>
//' Run C++ simulation
//' @param args command line arguments as a string vector
// [[Rcpp::export]]
Rcpp::CharacterVector
cpp_tumopp(const std::vector<std::string>& args) {
tumopp::Simulation simulation(args);
simulation.run();
return Rcpp::CharacterVector::create(
Rcpp::Named("config", simulation.config_string()),
Rcpp::Named("specimens", simulation.specimens()),
Rcpp::Named("drivers", simulation.drivers())
);
}
Try devtools::check().
Configure compile options
Error: R does not know where my C++ library is located:
run.cpp:3:10: fatal error: 'tumopp/simulation.hpp' file not found
#include <tumopp/simulation.hpp>
^~~~~~~~~~~~~~~~~~~~~~~
Write compile options to src/Makevars directly:
CXX_STD=CXX14
PKG_CPPFLAGS=-DSTRICT_R_HEADERS -I/usr/local/include
PKG_LIBS=-L/usr/local/lib -Wl,-rpath,/usr/local/lib -ltumopp
Or use configure/CMake script to generate it from src/Makevars.in:
CXX_STD=CXX14
PKG_CPPFLAGS=-DSTRICT_R_HEADERS @CPPFLAGS@
PKG_LIBS=@LDFLAGS@ @LDLIBS@
https://cran.r-project.org/doc/manuals/r-release/R-exts.html#Configure-and-cleanup
Transform C++ strings to data.frame
Create R/tumopp:
#' `tumopp()` returns full results with config columns in a data.frame
#' @param args command line arguments as a string vector
#' @export
tumopp = function(args = character(0L)) {
result = cpp_tumopp(c(0L, 0L, args))
population = readr::read_tsv(result["specimens"])
drivers = readr::read_tsv(result["drivers"])
readr::read_tsv(result["config"]) %>%
dplyr::mutate(population = list(population)) %>%
dplyr::mutate(drivers = list(drivers))
}
max coord dimensions shape delta0 rho0 population drivers
<int> <char> <int> <int> <num> <num> <list> <list>
1: 10000 hex 3 100 0.1 0.5 <tbl_df> <tbl_df>
Workflow improvement
Before using Rcpp:
- Run tumopp with some parameter sets
- Write results to TSV files
- Start R
- Read TSV files as data.frames
- Visualize and analyze
Thanks to Rcpp:
- Start R
- Run tumopp with some parameter sets in R
- Get results in a nested data.frame
- Visualize and analyze
Problem: Modern C++11/14/17 supported?
- Rcpp 0.12.10: C++17
- R 3.4.0: C++17
- OS and compiler
- macOS (clang): C++17
- Ubuntu 18.04 (gcc-7.3): C++17
- Ubuntu 16.04 (gcc-5.3): C++14
- CentOS 7 + Linuxbrew (gcc-5.5): C++14
- Windows + Rtools (gcc-4.9): C++11 ❌
- Windows + WSL + Ubuntu: ❓❓❓
http://gallery.rcpp.org/articles/rcpp-and-c++11-c++14-c++17/
🚧 Rtools 4 with gcc 8 is under development. (Thanks, Yutani-san!)
Tasks and Questions
✅ Core C++ library for tumor growth simulation
✅ R interface package using Rcpp and devtools
🚧 Visualization and analysis using tidyverse packages
🚧 Documentation using roxygen2, rmarkdown, and pkgdown
🚧 Tests using testthat
⬜ Hexagonal logo
✅ Publication doi:10.1371/journal.pone.0184229
✅ Advertisement in conferences (GSJ2017, SMBE2018, Tokyo.R#71)
⬜ Better way to transfer C++ data to R data.frames:
feather?
arrow?
⬜ C++ and R on Windows: WSL? Cygwin? MSYS? Docker?
✅ Talk over beers with YOU! 🍻
Reference
- https://github.com/heavywatal/tumopp
- https://github.com/heavywatal/rtumopp
- http://www.rcpp.org/
- http://adv-r.had.co.nz/Rcpp.html
- https://cran.r-project.org/doc/manuals/r-release/R-exts.html
- https://devtools.r-lib.org/
- https://usethis.r-lib.org/
- https://teuder.github.io/rcpp4everyone_ja/
- https://clang.llvm.org/cxx_status.html
- https://gcc.gnu.org/projects/cxx-status.html
- https://github.com/rwinlib/gcc-8